U-Net model for predicting cellular traction forces from fluorescence microscopy
I chose this project for my machine learning course project to understand the state of development in the machine learning applications to biophysics.
I had the following successful learning outcomes from this project:
- Understood the architecture of U-Net, one of the most successful neural net architecture in biomedical image segmentation (Ronneberger et al., 2015).
- Studied the work of (Schmidt et al., 2022) who used U-Net to predict traction forces from fluorescent images of cells.
- They create a dataset of fluorescent images of various proteins (zyxin, actin, paxillin, myosin) labels across multiple cells and corresponding measurement of traction forces using traction force microscopy.
- This way, they were able to train a U-Net to predict traction forces (both magnitude and direction) from fluorescent images.
- To test the generalizability of the Schimidt et al. model and get hands-on experience of concepts I learned during the course, I applied their model to images unseen by the model. I didn't perform any retraining to update the weights since I don't have access to a novel dataset. I just used the images I found in these publications which use TFM and label zyxin and actin: Kliewe et. al. 2024, Kudrayashov et al., 2022)
- One of the main things I found was that the binary mask threshold when marking the fluorescent parts of an image before feeding it to the model affects the output force prediction. For this project I set the threshold manually, by eyeballing the minimum required threshold to capture maximum information in a given image, but more rigourous analysis on this is required to be certain.
Here's an image showing images from Kliewe et. al. 2024, the corresponding binary masks for two different threshold values and the traction forces predicted by the model.
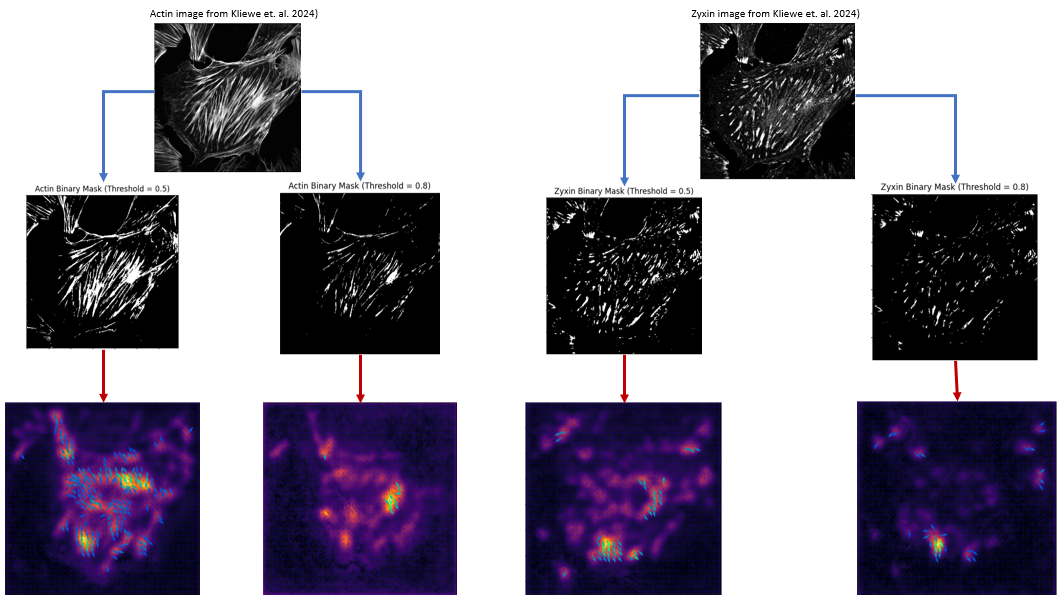 Traction force prediction from fluorescent images of cells using U-Net model. Top row images are from Kliewe et. al. 2024. Middle row images are binary masks created using two different threshold values. Bottom row images are corresponding traction forces predicted by Schmidt et al. U-Net model.
Traction force prediction from fluorescent images of cells using U-Net model. Top row images are from Kliewe et. al. 2024. Middle row images are binary masks created using two different threshold values. Bottom row images are corresponding traction forces predicted by Schmidt et al. U-Net model.While I don't have ground truth traction force data for these images to quantitatively assess the performance of the model, I am happy to get some kind of prediction that is not garbage. This shows that the model has some generalizability to unseen data, at least qualitatively.
Cytoskeleton dynamics
Biophysics
Machine learning
Python
Image analysis